The IEU GWAS Database
The MRC Integrative Epidemiology Unit (MRC IEU) at the University of Bristol hosts the IEU GWAS Database, one of the world’s largest open collections of Genome Wide Associate Study data. As of April 2019, the database contains over 250 billion genetic association records from more than 20,000 analysis of human traits.
The IEU GWAS database underpins the IEUs flagship MR-Base analytical platform (www.mrbase.org) which is used by people all over the world to carry out analyses that identify causal relationships between risk factors and diseases, and prioritize potential drug targets. The use of MR-Base by hundreds of unique users per week generates a high volume of queries to the IEU GWAS database (typically 1-4 million queries per week).
Objectives of the Oracle Cloud/MRC IEU collaboration
Both the IEU GWAS database and the MR-Base platform are open to the entire scientific community at no cost, supporting rapid knowledge discovery, and open and collaborative science. However, the scale of the database and the volume of use by academics, major pharma companies and others, create significant compute resource challenges for a non-profit organization. The primary objective of the collaboration was to explore ways to use Oracle Cloud Infrastructure to improve database performance and efficiency.
Methods
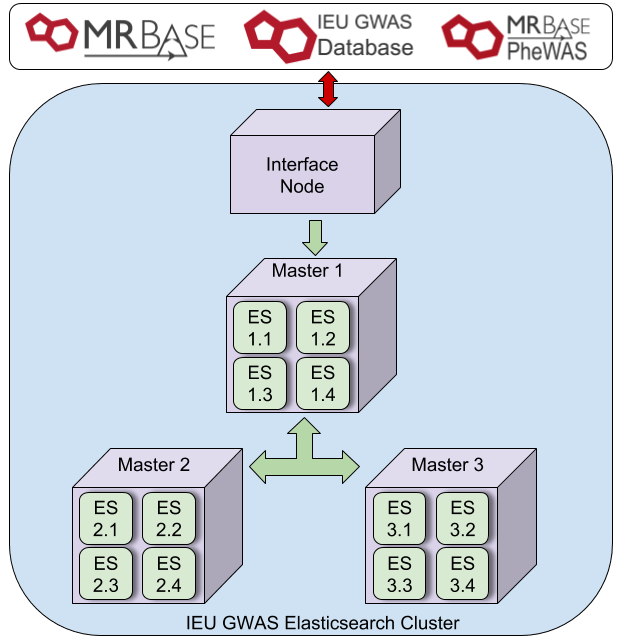
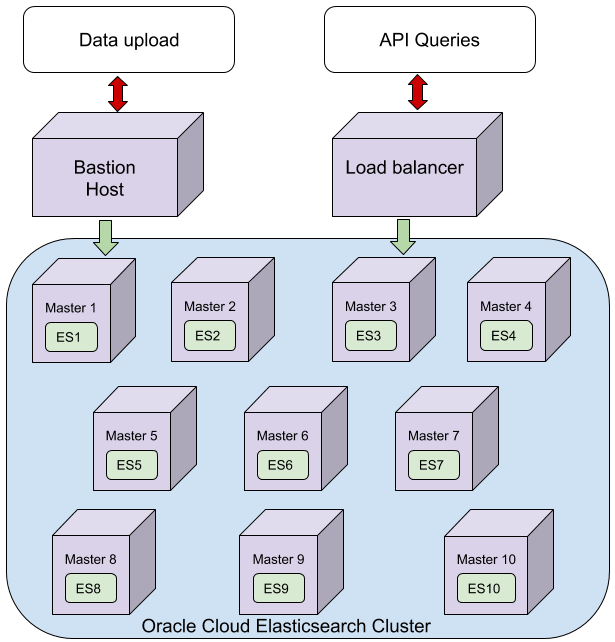
Previously, we have reported on our exiting solution for housing and searching our GWAS data. Our first configuration was to mirror our initial local setup, 3 machines with significant memory (384GB) and CPU (24). To replicate this in Oracle Cloud we modified an existing Elasticsearch repository (https://github.com/cloud-partners/oci-elastic) to match this, using 3 x VM.DenseIO2.24, each with 24 CPU, 320GB RAM and enough block storage to house the same amount of data. In addition we configured load balancers and a bastion node as per the original repo. The hardware here is significantly different to our local setup, with superior CPU, and SSD storage. After realising we didn’t actually need the block storage in this setup as each of the VMs contained 25.6 TB NVMe SSD (embarrassing!) our second setup was to use this onboard storage rather than the block storage. Thirdly, we configured 10x VM.Standard.E2.8 (8CPU, 64 GB RAM) with a similar load balancer, block storage and bastion node set up as configuration 1, to match (roughly) our current local architecture. In all three configurations a file store was used to provide NFS mounts across all nodes. A single snapshot of 500 GWAS was then transferred to the bastion node and restored to 40 separate indices, thus representing the full 200 billion UKBiobank records.
Testing
Elasticsearch is a search engine, great for full text indexing and super fast search, using focused queries that return a small number of records. Many of the queries we perform are indeed this type, single GWAS, single SNP, no problem. However, we do also perform larger, more demanding queries. For example, our PheWAS browser (http://phewas.mrbase.org/) searches for a single SNP across all records, extracting over 20,000 records at a time. This is not ideal for Elasticsearch, however performance locally was ok, around 30 seconds for an uncached query. Our most problematic queries involve multiple SNPS across all GWAS, filtered by a specific P-value. This could take a few minutes and although not a common query, something that represented the limit of our search capabilities. By performing 10 of these queries sequentially on random sets of SNPs, we could compare the configurations above. To check for caching we also performed 4 sequential tests.
Results
Performance

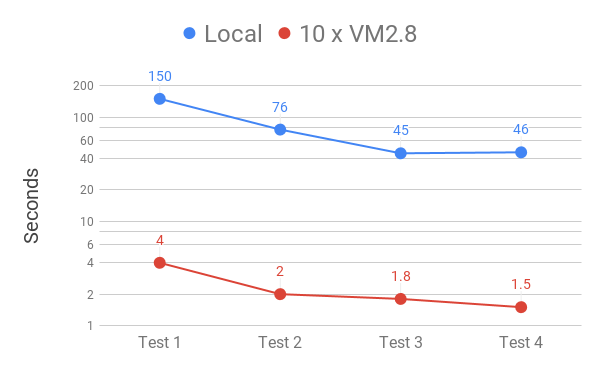
All three Oracle Cloud configurations were significantly quicker than our local instance. This was probably due to the SSD disk and better hardware. Interestingly the on-board NVME did not perform well, which was unexpected. The third configuration using 10 x VM.Standard.E2.8 performed best, with speeds almost 50 x quicker than our local cluster. This meant that a relatively complex query searching over 200 billion records could match and retrieve large numbers of records in just over a second! The PheWAS searches (not shown) were even quicker, with 10 in around 6 seconds.
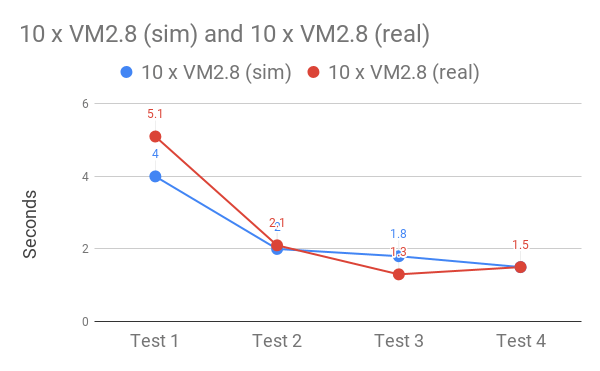
The final test was whether or not the performance improvements would hold when using real data. Fortunately, after much snapshotting, copying and indexing, a comparison of the simulated with real data revealed near identical query times.
Price
There is a cost to pay for this kind of performance. Pay as you go prices for this configuration would be around £40,000 per year, but estimates for up-front payment would be around half of this. Comparing these costs to local University infrastructure is difficult due to the mixture of direct and indirect costs, and factors such as the balance between hardware lifecycle and increasing data volume.
Discussion
One important question, is why do this, does it need to be that quick? We believe providing an API with this kind of search performance may change the way people use this kind of data. Research questions could become hypothesis free, instead of restricted to certain predefined areas. The ability to search data of this magnitude quickly and exhaustively could also be useful for drug development. Often, the time associated to a particular avenue of research is limited, and before all avenues have been explored it is time to move on. Providing the method to perform systematic checks across a large amount of data in a short space of time may ensure this isn’t an issue.
We plan to go live with the Oracle Cloud version of our database soon. This will give us time to test performance with real world queries and users, but we anticipate this development into the cloud will allow us to expand with our increasing numbers of GWAS and users.